Note
This page was generated from
Slices Alignment with Paste.ipynb.
Interactive online version:
.
Some tutorial content may look better in light mode.
Slices Alignment with PASTE#
This notebook demonstrates the process of Aligning spatial transcriptomics data. This is done in the following two steps:
Align spatial transcriptomics data from a set of multi-slices;
Reduce the amount of data by sampling to improve alignment speed.
Here we modified a published method PASTE, which utilized Fused Gromov-Wasserstein Optimal Transport (FGW-OT) algorithm, to leverages both gene expression similarity and spatial distances between spots to align and integrate spatial transcriptomics data.
Reference: Ron Zeira, Max Land, Benjamin J. Raphael. Alignment and Integration of Spatial Transcriptomics Data. bioRxiv, 2021.03.16.435604. doi: https://doi.org/10.1101/2021.03.16.435604
[1]:
import warnings
warnings.filterwarnings('ignore')
import spateo as st
2023-07-24 17:44:11.458267: W tensorflow/compiler/tf2tensorrt/utils/py_utils.cc:38] TF-TRT Warning: Could not find TensorRT
Load the data#
[2]:
# cellbin data
cellbin_slices = st.sample_data.drosophila(filename="E7-9h_cellbin_h5ad.zip")[4:8]
cellbin_slices
[2]:
[AnnData object with n_obs × n_vars = 1745 × 9057
obs: 'area', 'slices'
uns: '__type', 'spatial'
obsm: 'bbox', 'contour', 'spatial',
AnnData object with n_obs × n_vars = 1769 × 9158
obs: 'area', 'slices'
uns: '__type', 'spatial'
obsm: 'bbox', 'contour', 'spatial',
AnnData object with n_obs × n_vars = 1244 × 8992
obs: 'area', 'slices'
uns: '__type', 'spatial'
obsm: 'bbox', 'contour', 'spatial',
AnnData object with n_obs × n_vars = 1715 × 9145
obs: 'area', 'slices'
uns: '__type', 'spatial'
obsm: 'bbox', 'contour', 'spatial']
Align spatial transcriptomics data from a set of multi-slices#
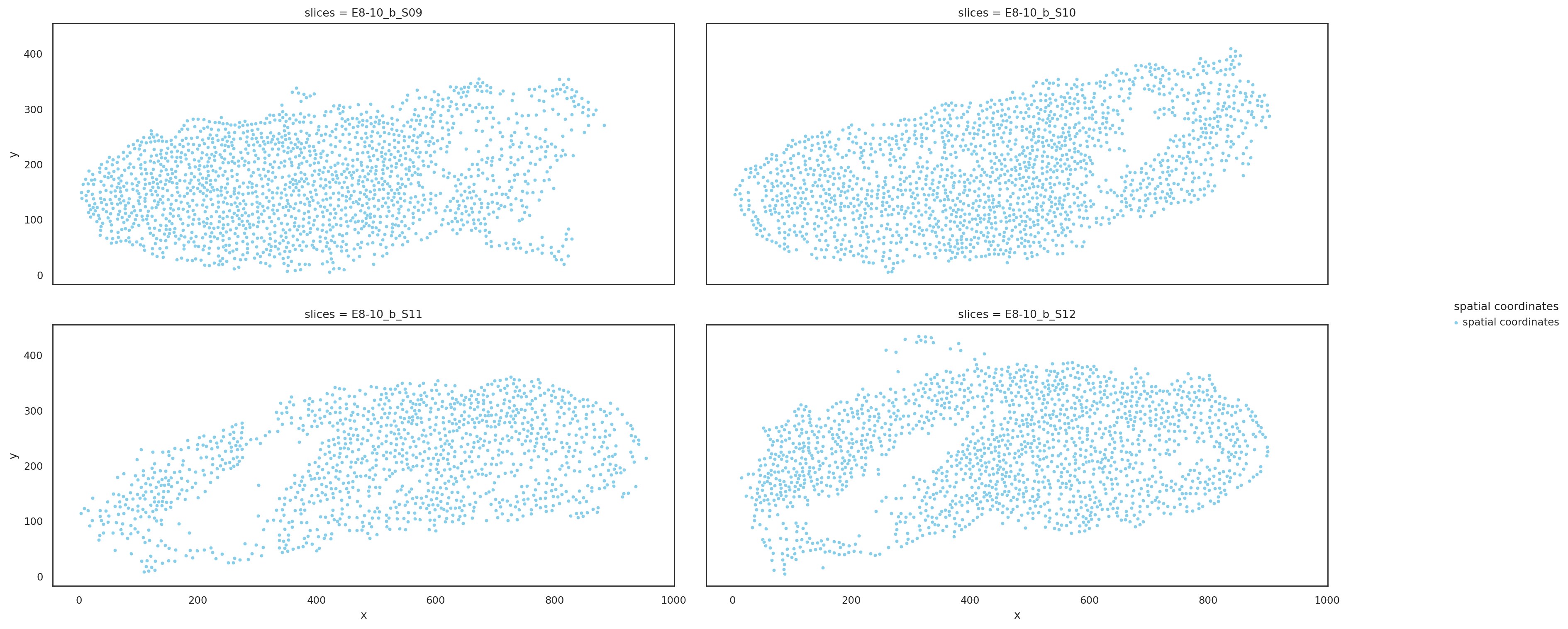
Visualize slices based on raw coordinates#
[3]:
st.pl.multi_slices(
slices=cellbin_slices,
slices_key="slices",
spatial_key="spatial",
point_size=10,
ncols=2,
)
Slices alignment#
[4]:
aligned_slices, pis = st.align.paste_align(
models=[slice.copy() for slice in cellbin_slices],
spatial_key="spatial",
key_added="align_spatial",
device="0" # or device="cpu"
)
|-----> [Models alignment] in progress: 33.3333%|-----> Filtered all samples for common genes. There are 8334 common genes.
|-----> [Models alignment] in progress: 66.6667%|-----> Filtered all samples for common genes. There are 8257 common genes.
|-----> [Models alignment] in progress: 100.0000%|-----> Filtered all samples for common genes. There are 8251 common genes.
|-----> [Models alignment] in progress: 100.0000%
|-----> [Models alignment] finished [13.8075s]
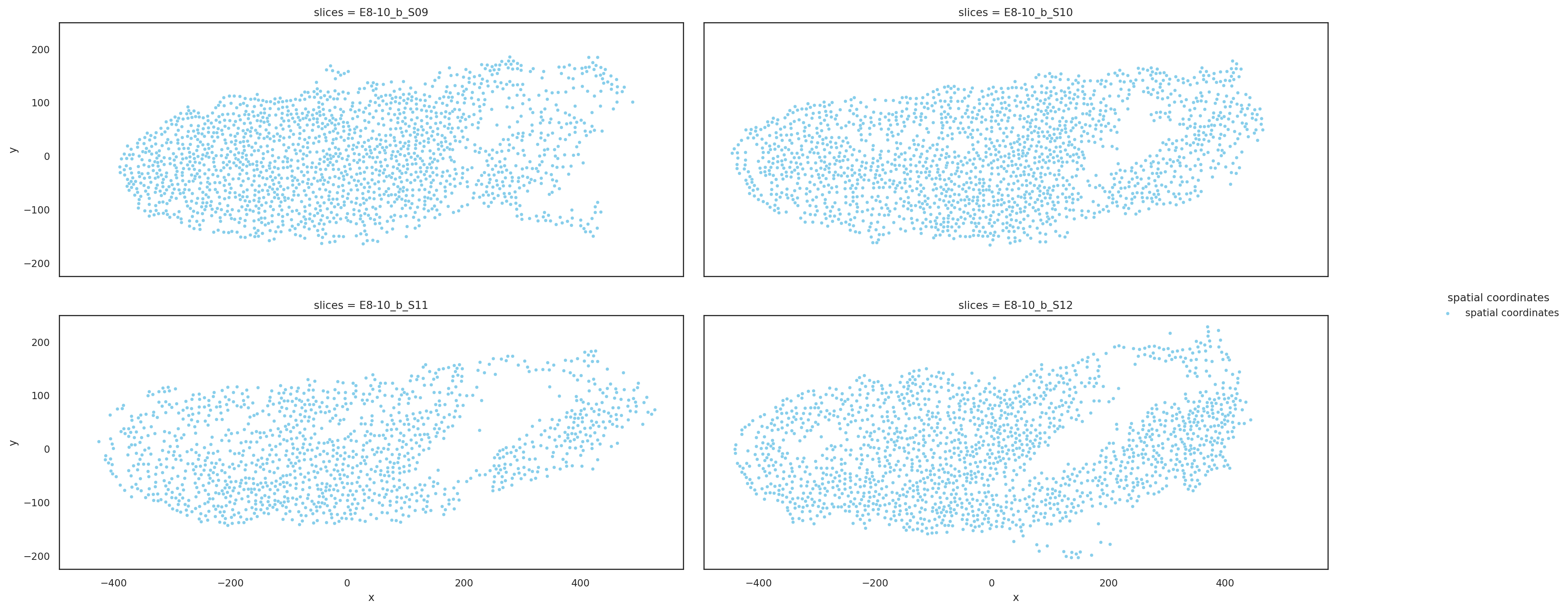
Visualize slices based on aligned coordinates#
[5]:
st.pl.multi_slices(
slices=aligned_slices.copy(),
slices_key="slices",
spatial_key="align_spatial",
point_size=10,
ncols=2,
)
Reduce the amount of data by down-sampling to improve alignment speed#
Slices alignment#
[6]:
aligned_slices, downsampling_slices, pis = st.align.paste_align_ref(
models=[slice.copy() for slice in cellbin_slices],
models_ref=None,
n_sampling=500,
sampling_method="trn",
spatial_key="spatial",
key_added="align_spatial",
numItermax=400,
device="0"
)
|-----> [Running TRN] in progress: 100.0000%|-----> [Running TRN] completed [19.5275s]
|-----> [Running TRN] in progress: 100.0000%|-----> [Running TRN] completed [19.2842s]
|-----> [Running TRN] in progress: 100.0000%|-----> [Running TRN] completed [19.7409s]
|-----> [Running TRN] in progress: 100.0000%|-----> [Running TRN] completed [19.7497s]
|-----> [Models alignment] in progress: 33.3333%|-----> Filtered all samples for common genes. There are 8334 common genes.
|-----> [Models alignment] in progress: 66.6667%|-----> Filtered all samples for common genes. There are 8257 common genes.
|-----> [Models alignment] in progress: 100.0000%|-----> Filtered all samples for common genes. There are 8251 common genes.
|-----> [Models alignment] in progress: 100.0000%
|-----> [Models alignment] finished [0.5662s]
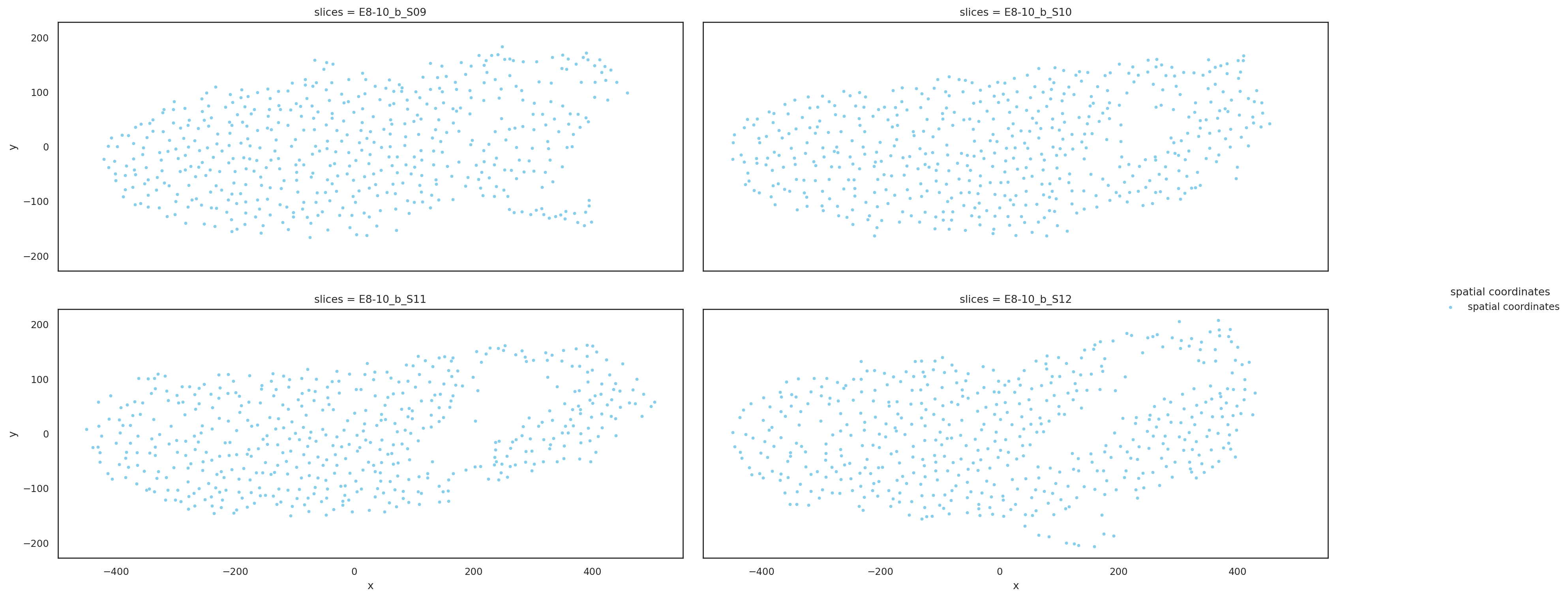
Visualize downsampling slices based on aligned coordinates#
[7]:
st.pl.multi_slices(
slices=downsampling_slices.copy(),
slices_key="slices",
spatial_key="align_spatial",
point_size=10,
ncols=2,
)
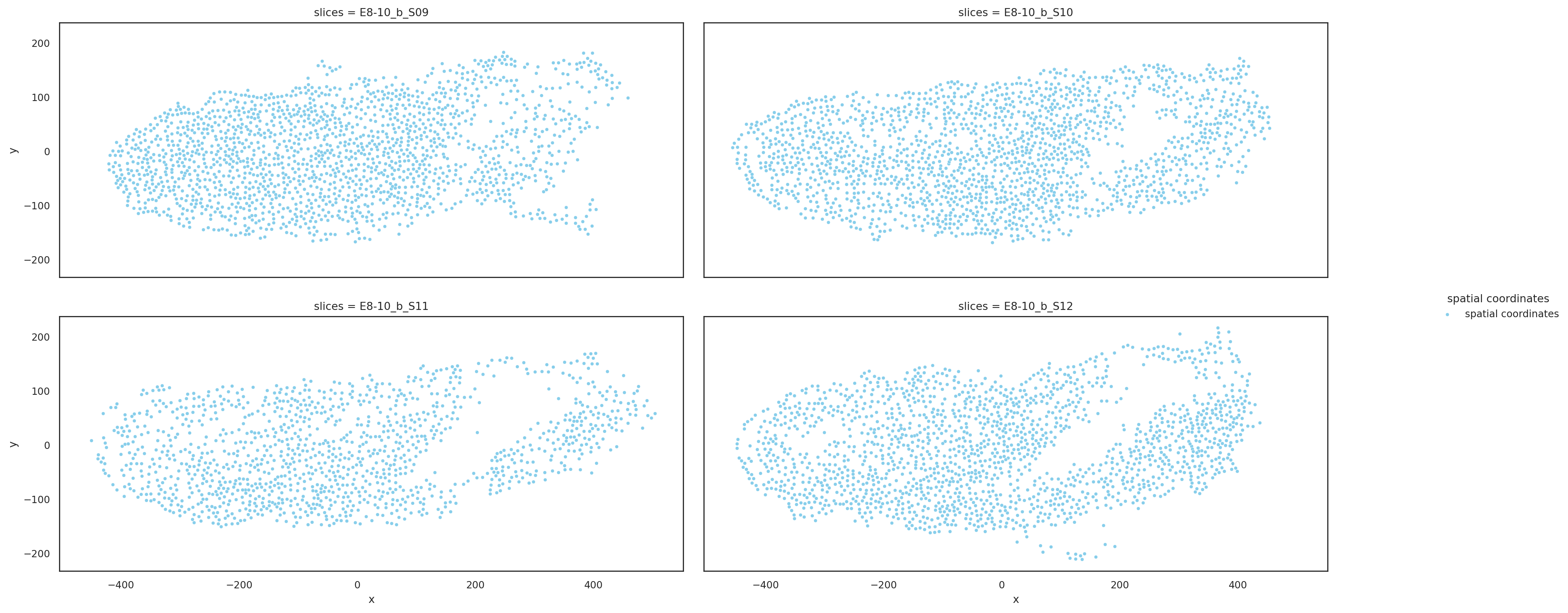
Visualize slices based on aligned coordinates#
[8]:
st.pl.multi_slices(
slices=aligned_slices.copy(),
slices_key="slices",
spatial_key="align_spatial",
point_size=10,
ncols=2,
)
[8]: